# -*- coding: utf-8 -*-
"""
author: 赵杭天
data: 20.1
base: official baseline
"""
from reader import data_loader
from glob import glob
from sklearn.cluster import KMeans
xmls_train = glob('./insects/train/annotations/xmls/*.xml')
xmls_val = glob('./insects/val/annotations/xmls/*.xml')
# print(len(xmls_train), len(xmls_val), len(xmls))
import xml.etree.ElementTree as ET
import os
import numpy as np
ct = 0
# records = [[{...}{...},...] , [{...},{...},...] ],第一个list放train的dict,第二个list放val的dict
records = [[],[]]
INSECT_NAMES = ['Boerner', 'Leconte', 'Linnaeus',
'acuminatus', 'armandi', 'coleoptera', 'linnaeus']
def get_insect_names():
"""
return a dict, as following,
{'Boerner': 0,
'Leconte': 1,
'Linnaeus': 2,
'acuminatus': 3,
'armandi': 4,
'coleoptera': 5,
'linnaeus': 6
}
It can map the insect name into an integer label.
"""
insect_category2id = {}
for i, item in enumerate(INSECT_NAMES):
insect_category2id[item] = i
return insect_category2id
for split_section, xmls in enumerate([xmls_train, xmls_val]):
cname2cid = get_insect_names()
for fpath in xmls:
tree = ET.parse(fpath)
if tree.find('id') is None:
im_id = np.array([ct])
else:
im_id = np.array([int(tree.find('id').text)])
objs = tree.findall('object')
im_w = float(tree.find('size').find('width').text)
im_h = float(tree.find('size').find('height').text)
gt_bbox = np.zeros((len(objs), 4), dtype=np.float32)
gt_class = np.zeros((len(objs),), dtype=np.int32)
is_crowd = np.zeros((len(objs),), dtype=np.int32)
difficult = np.zeros((len(objs),), dtype=np.int32)
for i, obj in enumerate(objs):
cname = obj.find('name').text
gt_class[i] = cname2cid[cname]
_difficult = int(obj.find('difficult').text)
x1 = float(obj.find('bndbox').find('xmin').text)
y1 = float(obj.find('bndbox').find('ymin').text)
x2 = float(obj.find('bndbox').find('xmax').text)
y2 = float(obj.find('bndbox').find('ymax').text)
x1 = max(0, x1)
y1 = max(0, y1)
x2 = min(im_w - 1, x2)
y2 = min(im_h - 1, y2)
# xywh
gt_bbox[i] = [(x1 + x2) / 2.0, (y1 + y2) / 2.0, x2 - x1 + 1., y2 - y1 + 1.]
is_crowd[i] = 0
difficult[i] = _difficult
voc_rec = {
'im_id': im_id,
'h': im_h,
'w': im_w,
'is_crowd': is_crowd,
'gt_class': gt_class,
'gt_bbox': gt_bbox,
'gt_poly': [],
'difficult': difficult
}
if len(objs) != 0:
records[split_section].append(voc_rec)
ct += 1
wh_total = [[],[]]
num_total = [[],[]]
for split_section, section_records in enumerate(records):
print('%s gt bbox数量:' % ('train' if split_section == 0 else 'val'))
# wh means width and height
wh_cls_wise = [[] for i in range(len(INSECT_NAMES))]
for r in section_records:
for i in range(len(r['gt_bbox'])):
bb = r['gt_bbox'][i][2:]
bb[0] = bb[0] / r['w']
bb[1] = bb[1] / r['h']
wh_total[split_section].append(bb)
wh_cls_wise[r['gt_class'][i]].append(bb)
for i, c in enumerate(wh_cls_wise):
print(INSECT_NAMES[i], ':\t', len(c))
num_total[split_section].append(len(c))
print('Total gt bbox: %d \n' % len(wh_total[split_section]))
# 绘图
import matplotlib.pyplot as plt
name_list = INSECT_NAMES
num_list_1 = num_total[0]
num_list_2 = num_total[1]
x =list(range(len(num_list_1)))
total_width, n = 0.8, 2
width = total_width / n
plt.figure(figsize=(10,8))
plt.bar(x, num_list_1, width=width, label='train', fc = 'g')
for i in range(len(x)):
x[i] = x[i] + width
plt.bar(x, num_list_2, width=width, label='val',tick_label = name_list,fc = 'y')
plt.legend()
# plt.show()
anchor_num = 9
kmeans = KMeans(n_clusters=anchor_num, random_state=0).fit(wh_total[0]+wh_total[1])
cluster_results = list(608 * kmeans.cluster_centers_)
cluster_results.sort(key=lambda wh: wh[1] * wh[0])
print('anchor_num = %d, 聚类结果:%s' % (anchor_num,cluster_results))
import matplotlib.patches as patches
plt.figure(2)
plt.imshow(np.full((100,100,3), 255))
currentAxis=plt.gca()
for bbox in cluster_results:
w, h = bbox
rect=patches.Rectangle((50-w/2, 50-h/2),w,h,linewidth=1,edgecolor='r',facecolor='none')
currentAxis.add_patch(rect)
plt.show()
各类别train/val统计柱状图
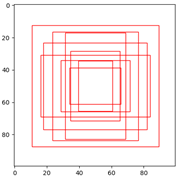
bbox聚类结果
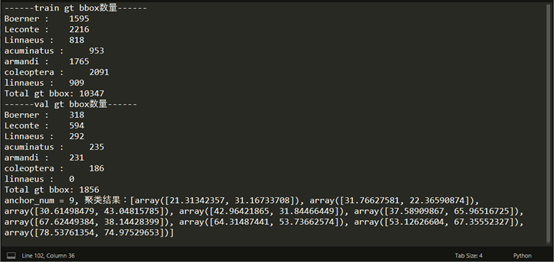
打印统计结果
留言